Our Mission
To understand how complex animal life evolved through changes in DNA and use this knowledge to become better stewards of the planet
The Genome 10K Community of Scientists: Assembling genomic data to understand vertebrate evolution and save dying species
Genome 10K is a project to sequence the genome of at least one individual from each vertebrate genus, approximately 10,000 genomes. It is a key milestone on the way toward the Vertebrate Genomes Project, the project to find and sequence at least one individual from each of the approximately 66,000 vertebrate species.
A mini-documentary about the Vertebrate Genomes Project
Scientists in the Vertebrate Genomes Project aim to sequence all vertebrates. That’s 66,000 species, maybe more. The goal: reference-grade sequence that helps labs around the world better understand these animals, their evolution, their ecosystems.
Toward a genome sequence for every animal: Where are we now?
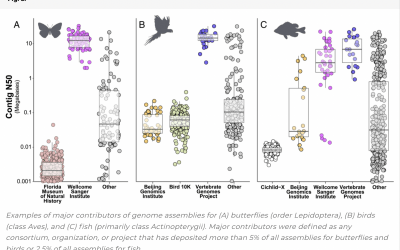
[Figure 5 from Article] Scott Hotaling, Joanna L. Kelley, and Paul B. Frandsen | PNAS | PNAS December 28, 2021 118 (52) e2109019118 In less than 25 y, the field of animal genome science has transformed from a discipline seeking its first glimpses into genome sequences...
Genomics-led solutions for all life on Earth
How will a landmark DNA sequencing project help secure the future of life – both ours and every other species'? In his talk at the 2021 Frontiers Forum Speaker Series, Professor Harris A. Lewin, American biologist, professor of evolution and ecology and Robert and...
Scientists describe ‘hidden biodiversity crisis’ as variation within species is lost
Scientists describe ‘hidden biodiversity crisis’ as variation within species is lost Many of the benefits people receive from nature depend on diversity within species, but this intraspecific variation is poorly understood and declining rapidly Tim Stephens | UCSC |...

Biodiversity Genomics 2020
Our annual G10K-VGP meeting, Biodiversity Genomics, was held October 5-9, 2020, celebrating success, exploring challenges and looking to the future of sequencing all life on Earth. The meeting was free to attend, thanks to generous support from the Tree of Life...